monocle3手动选取细胞发育轨迹分析
monocle3手动选取细胞发育轨迹分析
接之前分析:
monocle3相对于monocle2的升级我觉得最重要的就是可以手动在umap图赏选取起始细胞和终止细胞分析特定轨迹,这样可以个性化的分析自己关注的细胞亚群发育轨迹及关键表达变化的基因。
library(qs)
library(scop)
library(monocle3)
# 设置工作路径
setwd("~/maize_root/07.monocle3")
cds <- load_monocle_objects(directory_path = "monocle.Columella2//monocle3_cds.data/")
obj <- qread("../06.subset/Columella.added.subtype.qs") # 读入seurat对象
# 将cds UMAP替换seurat obj里面的umap
# 如果你跑monocle3是用的seurat降维结果(--use.seurat.umap),这步可以省略
umap_embeddings <- reducedDims(cds)$UMAP
colnames(umap_embeddings) <- c("umap_1", "umap_2")
obj[["umap"]]@cell.embeddings <- umap_embeddings[colnames(obj), ]
# 可视化结果
p1 <- plot_cells(cds,
color_cells_by = "subtype",
label_cell_groups = F,
label_groups_by_cluster = F,
label_leaves = T,
label_branch_points = TRUE,
reduction_method = "UMAP"
) +
ggtitle("trajectory") +
scale_color_manual(values = color_palette) +
theme(plot.title = element_text(hjust = 0.5))
p2 <- plot_cells(cds,
color_cells_by = "pseudotime",
label_cell_groups = FALSE,
label_leaves = FALSE,
label_branch_points = FALSE,
graph_label_size = 1.5
)
p1 + p2
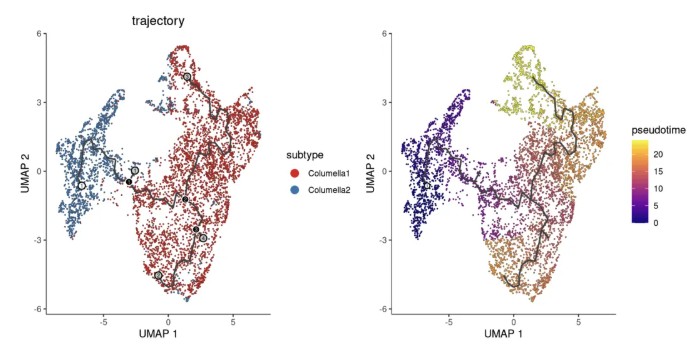
轨迹拐点ID显示:graph_label_size 点大小,label_principal_points 显示主点.
plot_cells(cds, color_cells_by = "pseudotime", label_cell_groups = FALSE, label_leaves = FALSE, label_branch_points = FALSE, label_principal_points = TRUE, group_label_size = 2, labels_per_group = 1, graph_label_size = 5 )
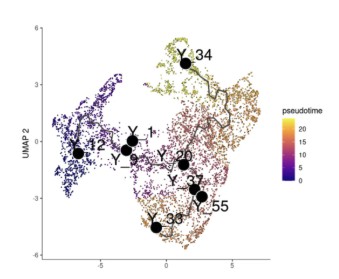
以Columella2 细胞为发育起点细胞,很明显有两个发育方向。根据节点名称选取感兴趣的两个细胞发育轨迹:
Y_12 -> Y_33
Y_12 -> Y_34
obj$Monocle3_Pseudotime <- pseudotime(cds)[colnames(obj)]
# Select the lineage using monocle3::choose_graph_segments
cds_sub <- choose_graph_segments(
cds,
starting_pr_node = "Y_12",
ending_pr_nodes = "Y_33"
)
obj$Lineage_1 <- NA
obj$Lineage_1[colnames(cds_sub)] <- obj$Monocle3_Pseudotime[colnames(cds_sub)]
CellDimPlot(
obj,
group.by = "subtype",
lineages = "Lineage_1",
lineages_span = 0.3,
theme_use = "theme_blank"
)
# 第二条发育轨迹
cds_sub <- choose_graph_segments(
cds,
starting_pr_node = "Y_12",
ending_pr_nodes = "Y_34"
)
obj$Lineage_2 <- NA
obj$Lineage_2[colnames(cds_sub)] <- obj$Monocle3_Pseudotime[colnames(cds_sub)]
CellDimPlot(
obj,
group.by = "subtype",
lineages = "Lineage_2",
lineages_span = 0.2,
theme_use = "theme_blank"
)
# 合并绘图
FeatureDimPlot(
obj,
features = "Monocle3_Pseudotime",
reduction = "UMAP",
lineages_span = 0.3,
lineages = paste0("Lineage_", 1:2),
palette = "inferno"
)
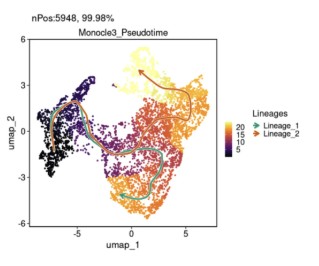
发育轨迹表达变化基因展示:
obj <- RunDynamicFeatures(
srt = obj,
lineages = c("Lineage_1", "Lineage_2"),
n_candidates = 200
)
ht <- DynamicHeatmap(
srt = obj,
lineages = c("Lineage_1", "Lineage_2"),
use_fitted = TRUE,
n_split = 6,
reverse_ht = "Lineage_1",
heatmap_palette = "viridis",
cell_annotation = "subtype",
pseudotime_label = 25,
pseudotime_label_color = "red",
height = 5,
width = 1.5
)
print(ht$plot)
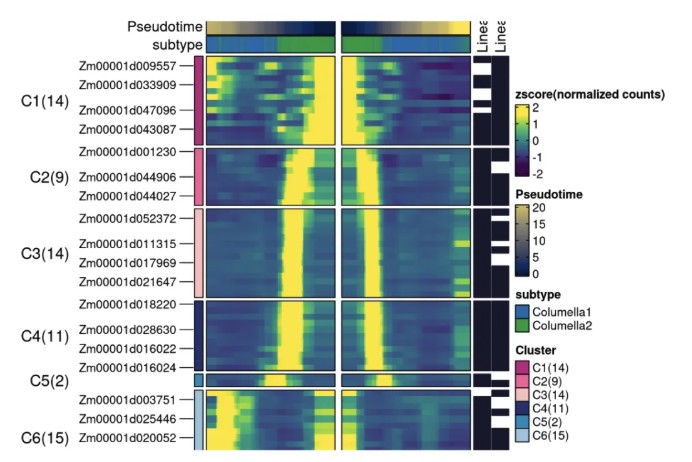
如果遇到软件安装错误无法运行,可以使用组学大讲堂云服务器,已经安装好所有单细胞转录组数据分析的软件运行环境,省去环境配置麻烦,详情可扫下方二维码咨询:

Wang, T., Wang, F., Deng, S. et al. Single-cell transcriptomes reveal spatiotemporal heat stress response in maize roots. Nat Commun16, 177 (2025). https://doi.org/10.1038/s41467-024-55485-3
跟多生信文章免费视频请关注,生信课堂:

- 发表于 2026-04-14 08:54
- 阅读 ( 343 )
- 分类:转录组
你可能感兴趣的文章
- 单细胞转录组拟时序-monocle3分析代码 402 浏览
- 单细胞拟时序分析monocle2-代码 586 浏览
- monocle3分析结果解读 12231 浏览
- Monocle2|单细胞测序的拟时序分析 10646 浏览
相关问题
- 老师,拟时序基础初步分析时,运行以下代码时Rscript $scripts/cell_trajectory_monocle3.r --rds ../04.cell_type_ann_leaf/leaf.added.celltype.qs \ --use.seurat.umap --groupby mycelltype -o monocle3.1 --cpu 6,出现图中报错,是因为聚类分群的时候resolution值太低了吗 2 回答
- 拟时分析筛选出的差异基因数量太少,有没有可以调整的参数去增加筛选数量 1 回答
- 单细胞测序使用subset脚本提出子集,再进行拟时分析报错plot_genes_by_group 1 回答
- 使用monocle3进行轨迹分析报错 2 回答
- monocle3 trajectory analysis 1 回答
0 条评论
请先 登录 后评论
