NCBI的SRA Lite和SRA Normalized数据有什么区别
有时候我们从NCBI下载SRA文件时会发现,会有SRA Lite和SRA Normalized两种类型的数据,前者后缀是.sralite,其文件大小也比原始的SRA文件要小。这两种数据的区别是什么呢?
ncbi下载SRR数据并转换为fastq数据:NCBI下载SRR并转换为fastq文件 - 组学大讲堂问答社区 (omicsclass.com)
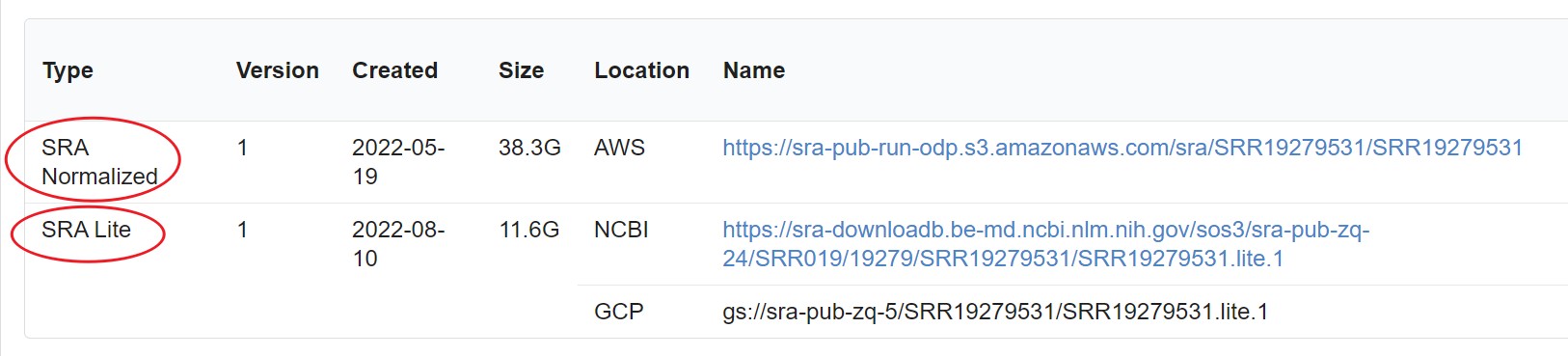
有时候我们从NCBI下载SRA文件时会发现,会有SRA Lite和SRA Normalized两种类型的数据,前者后缀是.sralite,其文件大小也比原始的SRA文件要小。这两种数据的区别是什么呢?
SRA Lite是NCBI里对SRA文件的另一个格式(https://www.ncbi.nlm.nih.gov/sra/docs/sra-data-formats/)
我们平常下载的SRA文件属于SRA Normalized Format,它和SRA Lite之间最大的区别就是SRA Lite采用了简化的quality scores,具体它是如何简化的请参考原文
SRA Lite files are produced from SRA Normalized Format by assessing overall read quality, setting a per-read quality flag (Read_Filter), and removing base quality scores from the file. In the resulting files, all reads have a Read_Filter flag with value pass or reject. Importantly, it is still possible to produce fastq formatted files from SRA Lite format using the SRA toolkit. In this case, each read will have a constant quality score set to 30 for reads with Read_Filter value "pass" or 3 for reads with a value "reject".
Illumina fastq and sam/bam specifications support a quality bit that is set by the sequencing instrument and SRA Lite stores this as a "pass"/"reject" Read_Filter value. If this bit is set in the submitted fastq or bam file, the value is retained. If it is not, SRA will set a pass/reject value based on the quality score distribution within each read. Reads that have more than half of quality score values <20 are flagged "reject". Reads that begin or end with a run of more than 10 quality scores <20 are also flagged "reject". Reads that pass these quality checks are flagged "pass". When dumping data using the fastq-dump, fasterq-dump, or sam-dump utilities in the SRA toolkit, all reads are included by default. However, the fastq-dump tool has an option to include only passed or only rejected reads:
fastq-dump --read-filter <[pass|reject]>
In order to interact with these files and set your preference for SRA Lite files, please use SRA Toolkit version 2.11.2 or later.
简单来说,SRA Lite文件将碱基质量得分分为了pass和reject两种,pass统一给分为30,而reject统一给分为3。这里我们要注意,在后面去接头等trim的分析步骤中,有些参数设置为去除碱基质量低于36的片段,在处理SRA Lite时要小心这一点。
那如何将SRA Lite转变为Fastq文件呢?做法是一样的,只需要保证SRA Toolkit软件的版本是最新的即可。还可以通过设置--read-filter参数为pass或reject来获得只包括pass或reject的Fastq文件。
参考:https://zhuanlan.zhihu.com/p/565413983
- 发表于 2023-06-08 14:29
- 阅读 ( 11091 )
- 分类:基础知识
你可能感兴趣的文章
- NCBI 10X 单细胞原始数据下载 解压注意:--include-technical 971 浏览
- 探索基因大数据的利器——SRA Explorer 4603 浏览
- NCBI 基因组文件下载——datasets 5397 浏览
- 使用NCBI——Batch Entrez批量下载蛋白序列 22640 浏览
- NCBI下载SRR并转换为fastq文件 18367 浏览
相关问题
- 请问老师怎么将NCBI里的gbff文件转化为gff格式的文件,因为里面找不到gff文件 0 回答
- 利用命令行进行sra数据下载有如下报错 1 回答
- NCBI序列上传 1 回答
- sra文件转换为fastq文件 1 回答
- NCBI序列上传 1 回答
- sratoolkit中prefetch下载一直报错是不是因为网的事情? 1 回答
0 条评论
请先 登录 后评论
