singleR细胞注释PBMC
singleR细胞注释
##调用第三方包
library(SingleR)
library(celldex)
library(qs)
library(Seurat)
##备份
setwd("~/pbmc/04.cell_type_ann/")
pbmc=qread("pbmc.added.celltype.qs")
pbmc_singleR <- pbmc
##载入参考数据集
MID <- MonacoImmuneData()
##提取表达矩阵
pbmc_count <- GetAssayData(object = pbmc_singleR,layer = "data")
##预测
pred <- SingleR(test = pbmc_count,
ref = MID,
assay.type.test=1,
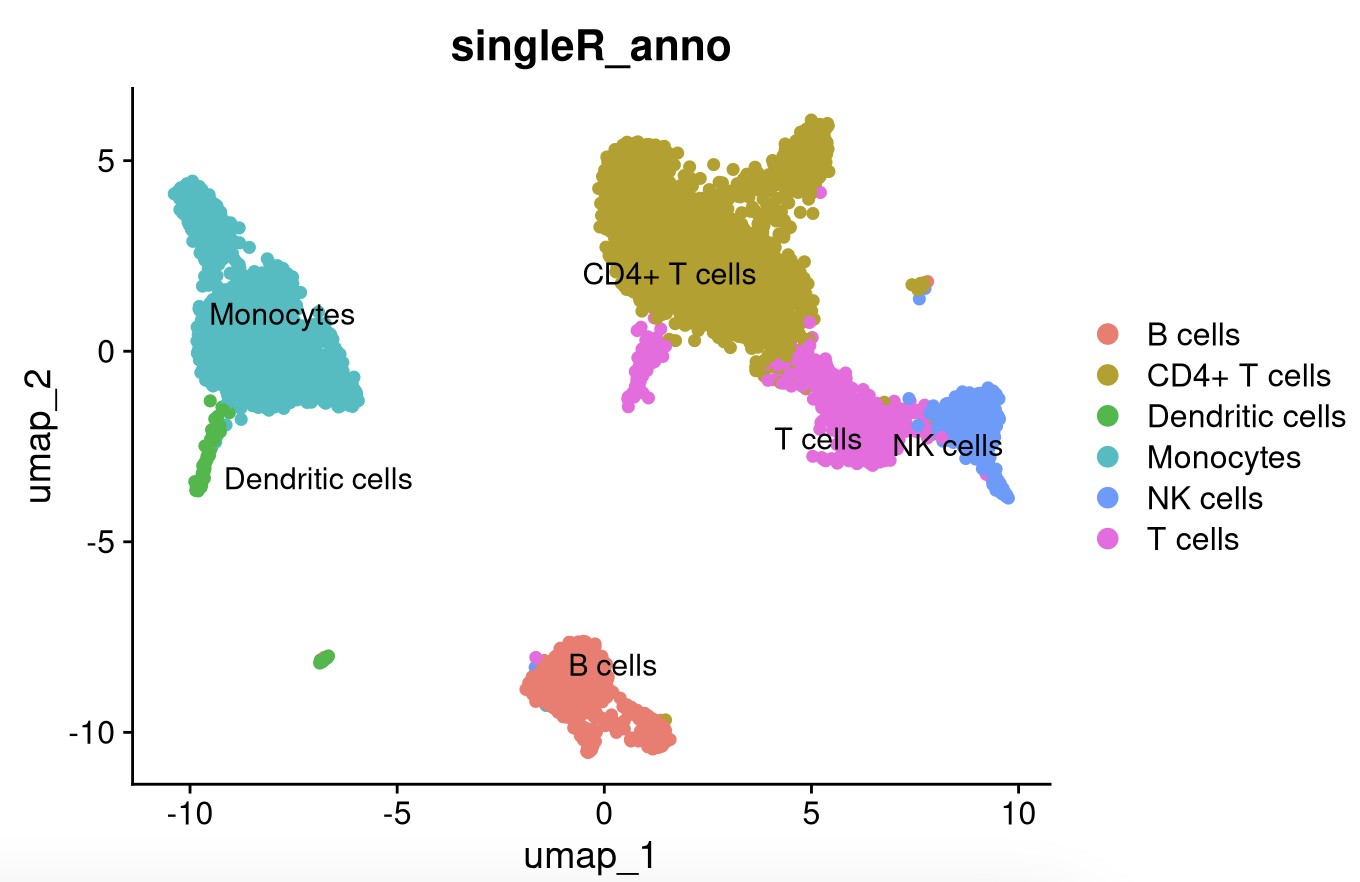
labels = MID$label.main,
clusters = pbmc_singleR$seurat_clusters)
pred
##提取并更新到pbmc_singleR中
pbmc_singleR$singleR_anno <- pred$labels[match(pbmc_singleR$seurat_clusters,rownames(pred))]
DimPlot(pbmc_singleR, reduction = "umap", label = TRUE, repel = TRUE,pt.size = 1.5,group.by = "singleR_anno")
pred <- SingleR(test = pbmc_count,
ref = MID,
assay.type.test=1,
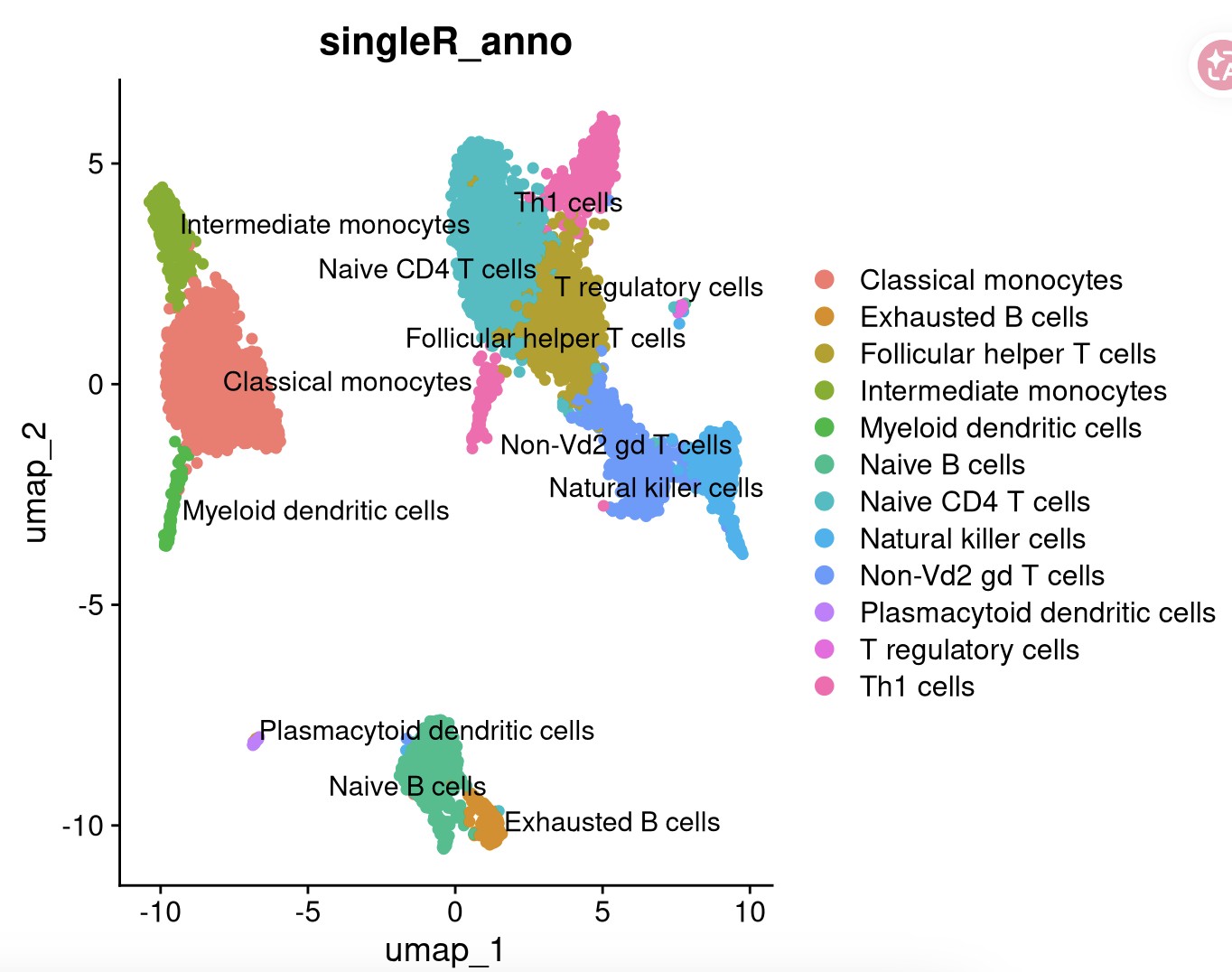
labels = MID$label.fine,
clusters = pbmc_singleR$seurat_clusters)
pred
##提取并更新到pbmc_singleR中
pbmc_singleR$singleR_anno <- pred$labels[match(pbmc_singleR$seurat_clusters,rownames(pred))]
DimPlot(pbmc_singleR, reduction = "umap", label = TRUE, repel = TRUE,pt.size = 1.5,group.by = "singleR_anno")
- 发表于 2026-04-15 15:40
- 阅读 ( 246 )
- 分类:转录组
你可能感兴趣的文章
相关问题
0 条评论
请先 登录 后评论
