热胁迫处理玉米根单细胞转录组差异基因分析
热胁迫处理玉米根单细胞转录组差异基因分析
热胁迫处理玉米根单细胞转录组差异基因分析
接之前分析:
Seurat R包提供三个方法查找差异基因,各自使用场景不同:
FindMarkers
用于进行精确的“一对一”比较,在用户指定的两个特定细胞群之间寻找差异基因,非常适合比较同一细胞类型在不同实验条件下的变化
FindAllMarkers
则是一种高效的“一对多”批量查找差异基因,它会自动为数据集中的每一个细胞群寻找相对于所有其他细胞的标记基因,是细胞类型注释和探索未知亚群身份时的标准流程;
FindConservedMarkers
专注于寻找“跨条件保守”的标记基因,它能在整合了多个样本或条件的数据集中,鉴定出在指定分组变量(如健康与疾病)里均稳定表达的保守标记,从而提供更可靠的跨样本生物标志物。
代码演示:
读入数据:
library(Seurat)
library("tidyverse")
library(qs)
library(scRNAtoolVis)
dimCols <- c(
"#E41A1C", "#377EB8", "#4DAF4A", "#984EA3", "#FF7F00", "#A65628",
"#F781BF", "#999999", "#66C2A5", "#FC8D62", "#8DA0CB", "#E78AC3",
"#A6D854", "#FFD92F", "#E5C494", "#B3B3B3", "#7FC97F", "#BEAED4",
"#FDC086", "#386CB0", "#F0027F", "#BF5B17", "#666666", "#1B9E77",
"#D95F02", "#7570B3", "#E7298A", "#66A61E", "#E6AB02", "#A6761D",
"#666666", "#A6CEE3", "#1F78B4", "#B2DF8A", "#33A02C", "#FB9A99",
"#E31A1C", "#FDBF6F", "#FF7F00", "#CAB2D6", "#6A3D9A", "#FFFF99",
"#B15928", "#8DD3C7", "#FFFFB3", "#BEBADA", "#FB8072", "#80B1D3",
"#FDB462", "#B3DE69", "#FCCDE5", "#D9D9D9", "#BC80BD", "#CCEBC5",
"#FFED6F", "#FBB4AE", "#B3CDE3", "#CCEBC5", "#DECBE4", "#FED9A6",
"#FFFFCC", "#E5D8BD", "#FDDAEC", "#F2F2F2", "#B3E2CD", "#FDCDAC",
"#CBD5E8", "#F4CAE4", "#E6F5C9", "#FFF2AE", "#F1E2CC", "#CCCCCC"
)
# 读入前面聚类结果seurat对象qs文件
obj <- qread(
"~/maize_root/04.cell_type_ann/root.added.celltype.qs",
nthreads = 4
)
# 常用参数:only.pos = TRUE 非常常用,
# 它会让结果只返回在该群中高表达(阳性)的标记基因
FindAllMarkers: 查找细胞亚群高表达基因
markers <- FindAllMarkers(obj,
group.by = "celltype",
logfc.threshold = 0.1,
test.use = "wilcox",
only.pos = FALSE)
#过滤一下
markers %>%
group_by(cluster) %>%
dplyr::filter(abs(avg_log2FC) > 1)
#热图top5基因展示
markers %>%
group_by(cluster) %>%
dplyr::filter(avg_log2FC > 1) %>%
slice_head(n = 5) %>%
ungroup() -> top5
DoHeatmap(obj, features = top5$gene,group.colors=dimCols) +
NoLegend()
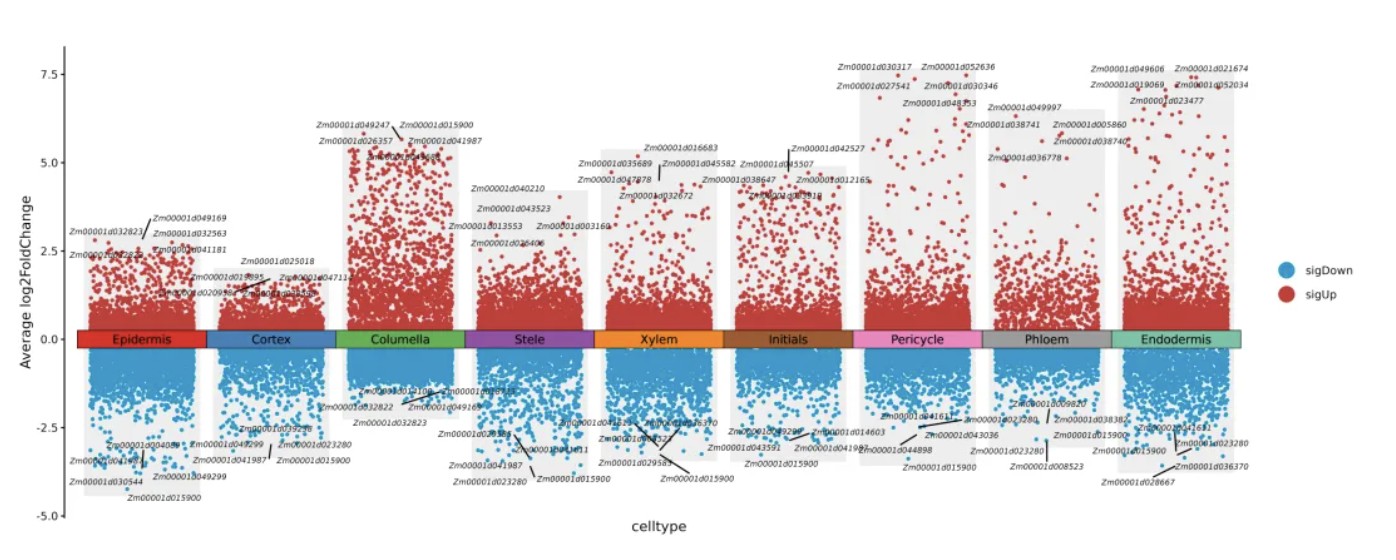
差异基因可视化火山图
jjVolcano(diffData = markers,
log2FC.cutoff= 1,
pvalue.cutoff= 0.05,
adjustP.cutoff= 0.001,
pSize=0.5,
size=2,
legend.position = NULL,
tile.col=dimCols)
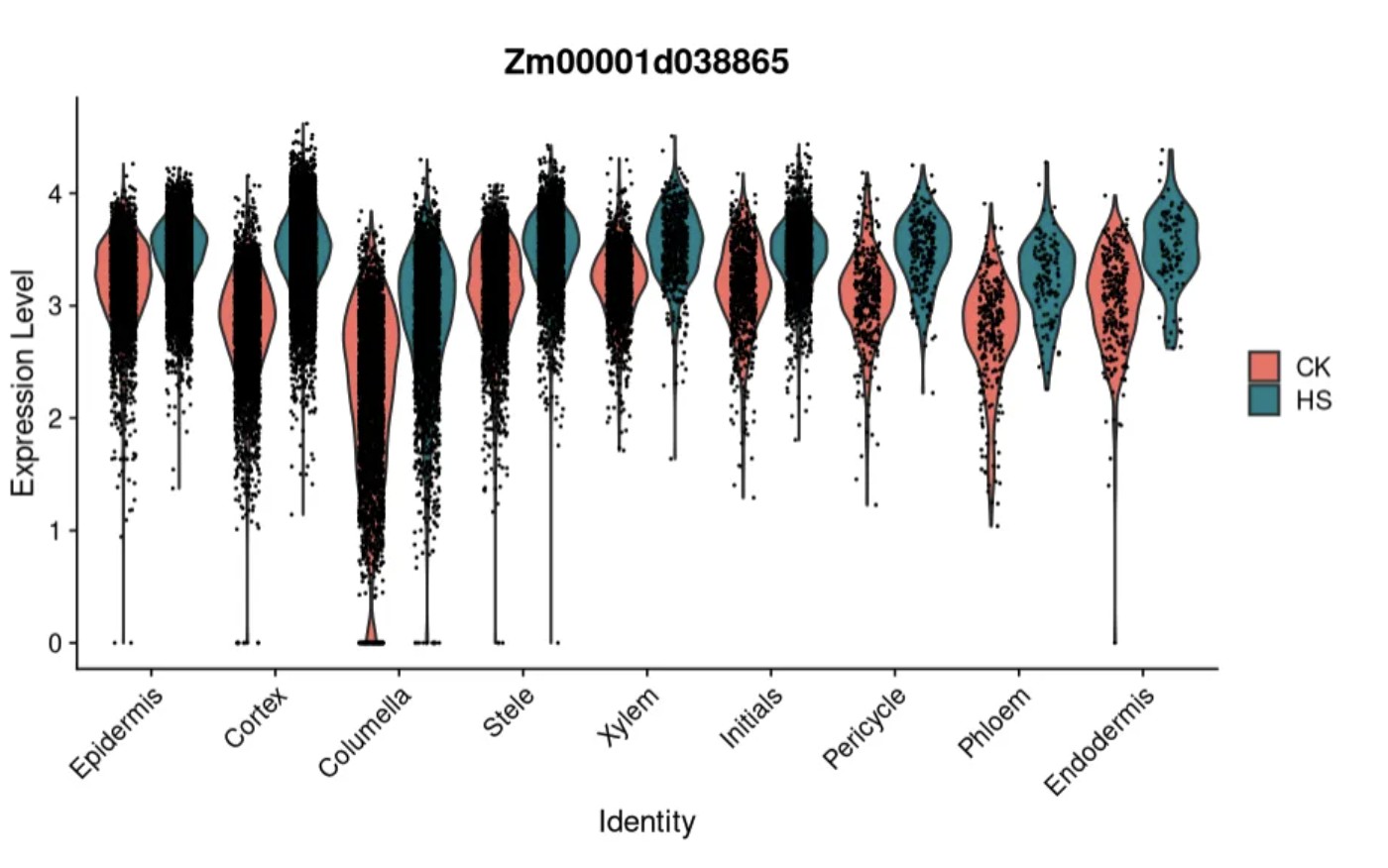
FindMarkers:Cortex细胞热处理后高表达基因,热响应基因:
Cortex.degs<- FindMarkers(obj, ident.1 = "HS", ident.2 = "CK",
group.by="Group",
subset.ident="Cortex",
logfc.threshold = 0.25,
test.use = "wilcox", only.pos = TRUE)
VlnPlot(obj,features = "Zm00001d038865",group.by = "celltype",split.by = "Group")
这样可以找到Cortex细胞热处理前后差异表达基因:
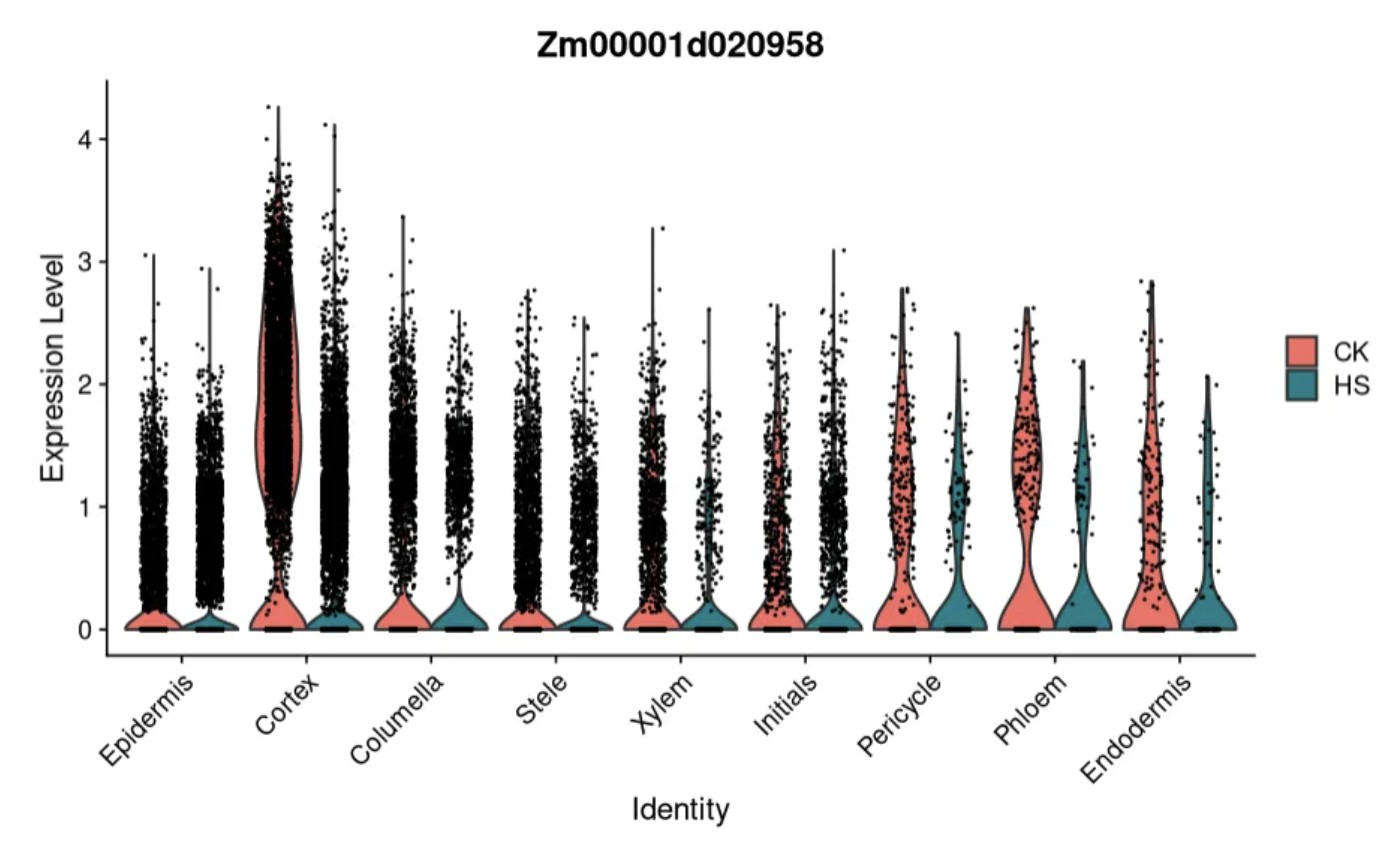
FindConservedMarkers:热处理前后cortex细胞中保守的基因,也就是不受热处理影响的基因:
Cortex.conserved<- FindConservedMarkers(obj, ident.1 = "Cortex", ident.2 = NULL,
grouping.var="Group",
logfc.threshold = 0.25,
test.use = "wilcox", only.pos = TRUE)
VlnPlot(obj,features = "Zm00001d020958",group.by = "celltype",split.by = "Group")
如果遇到软件安装错误无法运行,可以使用组学大讲堂云服务器,已经安装好所有单细胞转录组数据分析的软件运行环境,省去环境配置麻烦,详情可扫下方二维码咨询:

- 发表于 2026-04-08 15:24
- 阅读 ( 286 )
- 分类:转录组
你可能感兴趣的文章
相关问题
0 条评论
请先 登录 后评论
