可变剪切绘图 | rmats2sashimiplot
在真核生物里,基因转录出来的是 pre-mRNA(前体 mRNA),里面包含外显子和内含子。在变成成熟 mRNA 的过程中,内含子会被切掉,外显子会被拼接起来,这个过程就叫 剪切(splicing)。而可变剪切(Alternative Splicing, AS) 就是:同一个基因,在不同条件、不同组织、不同处理下,可以选择不同的外显子组合,拼接成多条不同的 mRNA,最终翻译成不同的蛋白质。通常在转录组分析中,会进行可变剪切的分析。
1.可变剪切基本类型
rMATS 最经典的就是把可变剪切分成 5 种基本模式:
1.SE:外显子跳跃(Skipped Exon)某个外显子直接被跳过,不包含在最终 mRNA 里。
2.RI:内含子保留(Retained Intron)本该切掉的内含子被保留下来。
3.A5SS:可变 5’ 剪切位点(Alternative 5' Splice Site)同一个外显子,5’端有两个剪切位点可选。
4.A3SS:可变 3’ 剪切位点(Alternative 3' Splice Site)同一个外显子,3’端有两个剪切位点可选。
5.MXE:互斥外显子(Mutually Exclusive Exons)两个相邻外显子,永远只出现一个,不会同时出现。
这 5 种基本类型,基本覆盖了绝大多数可变剪切事件。
2.分析流程
rMATS是目前转录组可变剪切分析最常用、最稳定的软件之一,专门用于两组样本之间的可变剪切差异分析
1.rMATS
python rmats.py --b1 A.txt --b2 B.txt \ --gtf ref.gtf --nthread 线程数 --od 输出文件夹名称 \ -t paired --variable-read-length \ --readLength 150 --cstat 0.0001 \ --libType fr-unstranded --novelSS
其中A.txt和B.txt里面是A组和B组样本的bam文件位置:
A.txt:a1.bam,a2.bam,a3.bam
B.txt:b1.bam,b2.bam,b3.bam
其他参数如下:
python rmats/bin/rmats.py -h
usage: rmats.py [options]
options:
-h, --help show this help message and exit
--version show program's version number and exit
--gtf GTF An annotation of genes and transcripts in GTF format
--b1 B1 A text file containing a comma separated list of the BAM files for sample_1. (Only if using BAM)
--b2 B2 A text file containing a comma separated list of the BAM files for sample_2. (Only if using BAM)
--s1 S1 A text file containing a comma separated list of the FASTQ files for sample_1. If using paired reads the format is ":" to separate pairs and "," to separate replicates. (Only if using fastq)
--s2 S2 A text file containing a comma separated list of the FASTQ files for sample_2. If using paired reads the format is ":" to separate pairs and "," to separate replicates. (Only if using fastq)
--od OD The directory for final output from the post step
--tmp TMP The directory for intermediate output such as ".rmats" files from the prep step
-t {paired,single} Type of read used in the analysis: either "paired" for paired-end data or "single" for single-end data. Default: paired
--libType {fr-unstranded,fr-firststrand,fr-secondstrand} Library type. Use fr-firststrand or fr-secondstrand for strand-specific data. Only relevant to the prep step, not the post step. Default: fr-unstranded
--readLength READLENGTH .The length of each read
--variable-read-length Allow reads with lengths that differ from --readLength to be processed. --readLength will still be used to determine IncFormLen and SkipFormLen
--anchorLength ANCHORLENGTH . The "anchor length" or "overhang length" used when counting the number of reads spanning splice junctions. A minimum number of "anchor length" nucleotides must be mapped to each end of a given junction. The minimum value is 1 and the default value is set to 1 to make use of all possible splice junction reads.
--tophatAnchor TOPHATANCHOR. The "anchor length" or "overhang length" used in the aligner. At least "anchor length" NT must be mapped to each end of a given junction. The default is 1. (Only if using fastq)
--bi BINDEX The directory name of the STAR binary indices (name of the directory that contains the SA file). (Only if using fastq)
--nthread NTHREAD The number of threads. The optimal number of threads should be equal to the number of CPU cores. Default: 1
--tstat TSTAT The number of threads for the statistical model. If not set then the value of --nthread is used
--cstat CSTAT The cutoff splicing difference. The cutoff used in the null hypothesis test for differential splicing. The default is 0.0001 for 0.01% difference. Valid: 0 <= cutoff < 1. Does not apply to the paired stats model
--task {prep,post,both,inte,stat}
Specify which step(s) of rMATS to run. Default: both. prep: preprocess BAMs and generate a .rmats file. post: load .rmats file(s) nto memory, detect and count alternative splicing events, and calculate P value (if not --statoff). both: prep + post. inte(integrity): check that the BAM filenames recorded by the prep task(s) match the BAM filenames for the current command line. stat: run statistical test on existing output files
--statoff Skip the statistical analysis
--paired-stats Use the paired stats model
--novelSS Enable detection of novel splice sites (unannotated splice sites). Default is no detection of novel splice sites
--mil MIL Minimum Intron Length. Only impacts --novelSS behavior. Default: 50
--mel MEL Maximum Exon Length. Only impacts --novelSS behavior. Default: 500
--allow-clipping Allow alignments with soft or hard clipping to be used
--fixed-event-set FIXED_EVENT_SET A directory containing fromGTF.[AS].txt files to be used instead of detecting a new set of events
最后得到的结果如下:
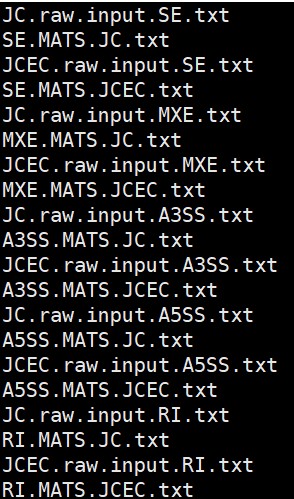
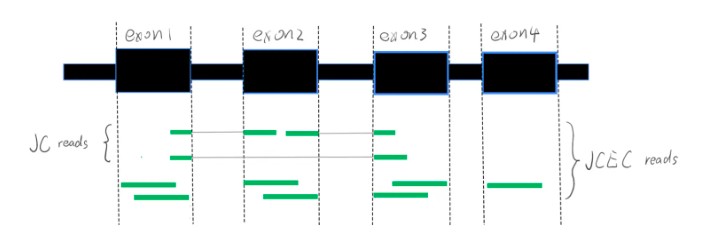
rMATS中,JC是JunctionCounts的缩写,表示跨越剪切位点的reads数量。JCEC是JunctionCounts和ExonCounts的缩写合并,Exon Counts表示不跨越剪切位点的reads数量,JCEC可以理解为所有比对上的reads。
其中一个重要结果SE.MATS.JC.txt结果:

2.rmats2sashimiplot绘图
2.1 rmats2sashimiplot下载安装
git clone https://gitcode.com/gh_mirrors/rm/rmats2sashimiplot
cd rmats2sashimiplotpython2 setup.py install
# 可以提前安装需要的依赖:
pip install numpy scipy matplotlib pysam
# 如果安装不成功怎么使用:
#rmats2sasmimiplot can be run without installing:
python ./src/rmats2sashimiplot/rmats2sashimiplot.py
安装好了之后存在很多问题,主要是库导入的问题,还有python2到3的一个问题,建议使用python2运行,然后手动修改部分脚本解决库导入。
2.2 rmats2sashimiplot绘图
python2 /share/work/biosoft/rmats2sashimiplot/rmats2sashimiplot/src/rmats2sashimiplot/rmats2sashimiplot.py \
--b1 a1.bam,a2.bam,a3.bam \
--b2 b1.bam,b2.bam,b3.bam \
--event-type SE -e SE.MATS.JC.txt \
--l1 One --l2 Two --exon_s 1 --intron_s 5 -o test_events_output
这里用到的是bam文件所以输入文件的参数是--b1和--b2,如果是sam文件记得改成--s1和--s2。其他参数解释如下
--event-type 事件类型,从01里面的五个类型选择,也可以根据*.JC.txt的前缀写
-e rMATs结果文件
-l1 第一个组的lable
-l2 第二个组的lable
--exon_s How much to scale down exons.
--intron_s How much to scale down introns
2.3 结果展示
结果位于./test_events_output/Sashimi_plot文件夹里面
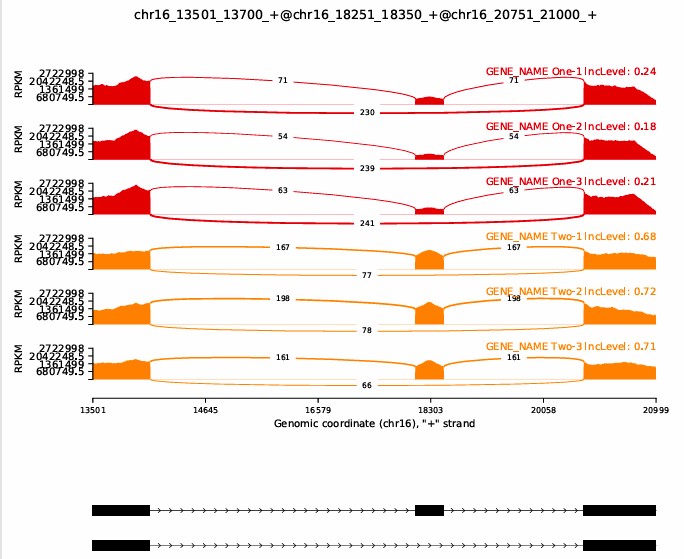
References:
https://www.jianshu.com/p/d09b95a98c64
https://github.com/Xinglab/rmats2sashimiplot
https://github.com/Xinglab/rmats-turbo
- 发表于 2026-02-27 12:00
- 阅读 ( 451 )
- 分类:转录组
你可能感兴趣的文章
- 可变剪切在癌症中的研究 6304 浏览
