华大DNBC4单细胞转录组数据分析-拟南芥根愈伤组织单细胞转录组图谱

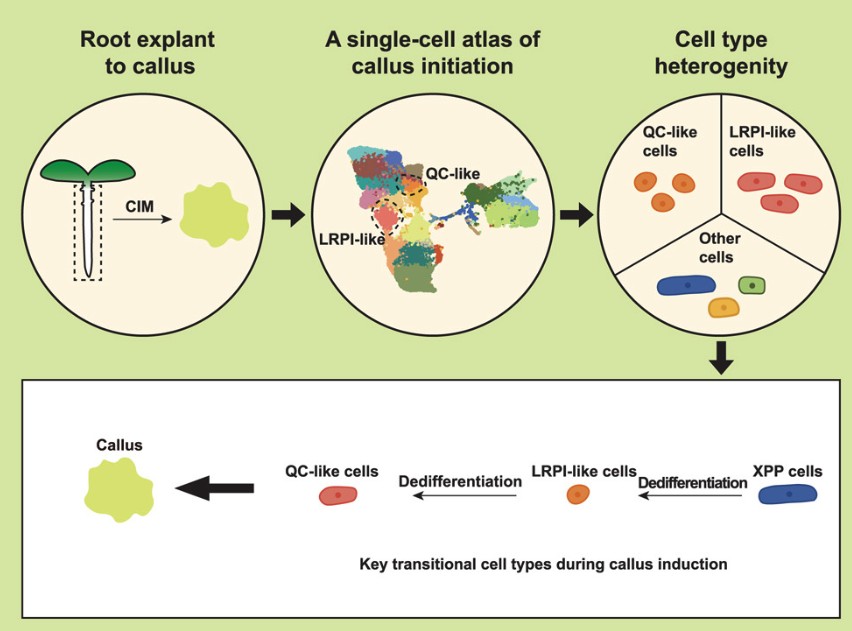
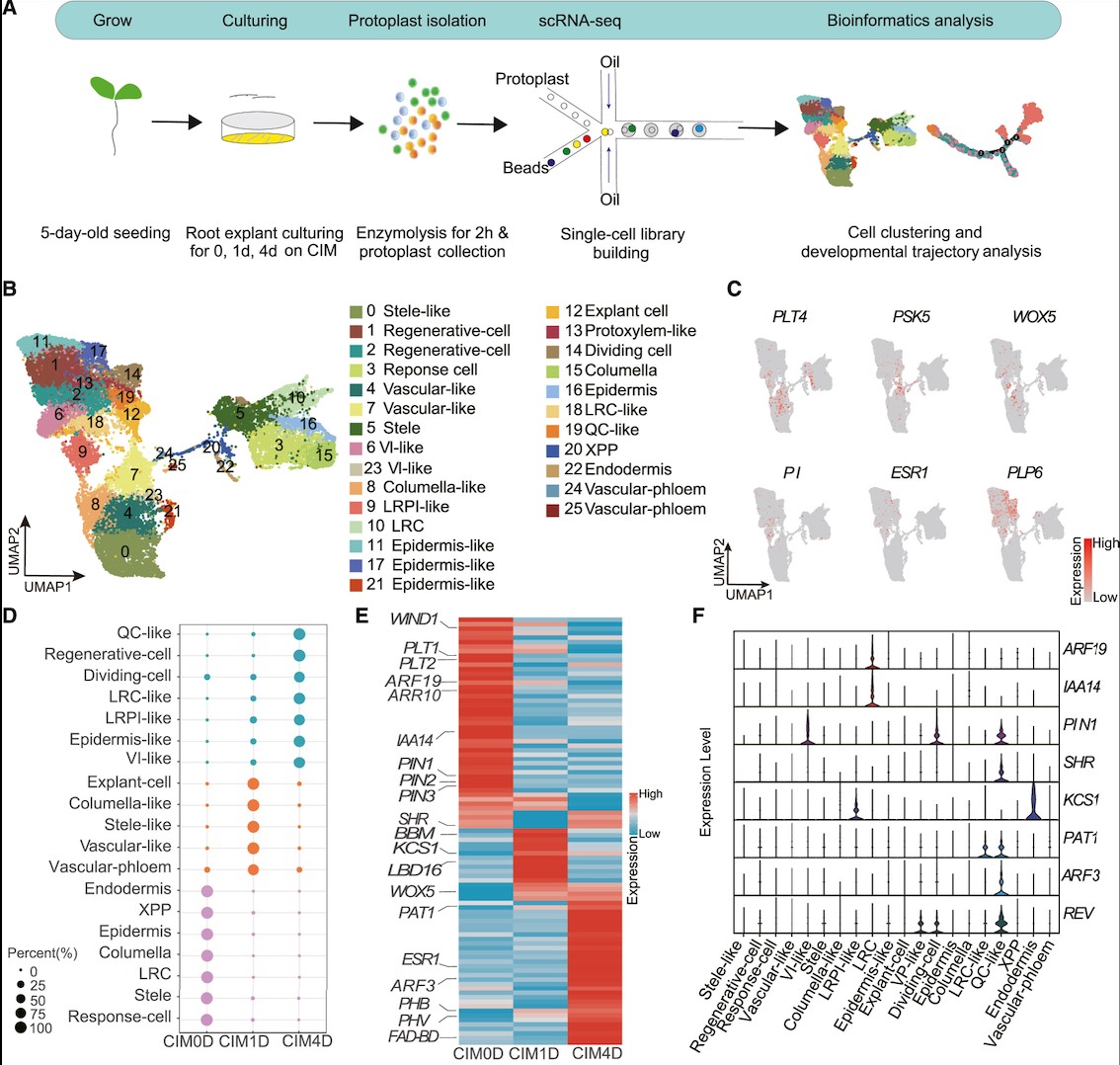
Yin 等人(2024)利用时间序列单细胞转录组技术,对拟南芥根外植体在愈伤诱导培养基(CIM)上培养 0、1、4 天的样本进行解析,构建了愈伤发生早期的高精度单细胞图谱,总细胞数达 25188 个,成功注释出 26 个细胞簇与多种细胞类型。实验设计以时间序列为核心,追踪从根体细胞到愈伤细胞的命运转变,取样为 5 日龄拟南芥幼苗根外植体,经酶解制备原生质体后进行 scRNA‑seq,结合聚类、拟时轨迹、CytoTRACE 分化潜能评估与转录因子调控网络(TFRN)分析,系统解析细胞去分化与多能性获得过程。研究发现,木质部极中柱鞘(XPP)细胞是愈伤起始的源头细胞,经诱导形成LRPI‑like 细胞(类侧根原基起始细胞),该类细胞作为关键中间态,高表达侧根起始与脂质转运相关基因(KCS6、LTP1、LTP2),并进一步脱分化形成QC‑like 细胞;QC‑like 细胞具有最低分化程度与最高多能性得分,特异高表达再生核心基因 ESR1、PHB、PHV、WOX5,是愈伤组织中核心多能干细胞群。拟时分析证实,LRPI‑like 细胞沿假时间轨迹向 QC‑like 细胞转变,该过程伴随侧根发育基因(LBD16、PLT4)下调与愈伤多能性基因上调;同时生长素信号、细胞周期与细胞壁重塑基因持续激活,支撑细胞命运转换。转录因子调控网络分析鉴定出 16 个核心转录因子(如 TCP23、RTV1、bHLH104、WOX13)与 106 个靶基因,调控细胞分裂、细胞壁合成与分生组织发育,其中 PHB、REV、CYCD3;3 为关键功能基因。该研究明确了愈伤发生的核心轨迹:XPP → LRPI‑like → QC‑like,揭示了细胞从体细胞逐步去分化获得多能性的分子机制,同时证实愈伤形成与侧根发育路径不同,为理解植物细胞全能性与优化遗传转化再生体系提供了关键理论与靶点。
数据下载链接:
https://db.cngb.org/data_resources/project/CNP0004389
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809366/CNX0734939/CNR0835613/FP200003884_L01_31_2.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809365/CNX0734938/CNR0835612/FP200003884_L01_27_2.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809368/CNX0734941/CNR0835615/DP8400024433BL_L01_80_2.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809367/CNX0734940/CNR0835614/DP8400024433BL_L01_78_2.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809361/CNX0734936/CNR0835610/DP8400025960BR_L01_97_1.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809366/CNX0734939/CNR0835613/FP200003884_L01_31_1.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809363/CNX0734937/CNR0835611/DP8400026128BL_L01_71_1.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809363/CNX0734937/CNR0835611/DP8400026128BL_L01_71_2.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809368/CNX0734941/CNR0835615/DP8400024433BL_L01_80_1.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809365/CNX0734938/CNR0835612/FP200003884_L01_27_1.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809367/CNX0734940/CNR0835614/DP8400024433BL_L01_78_1.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data5/CNP0004389/CNS0809361/CNX0734936/CNR0835610/DP8400025960BR_L01_97_2.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data2/CNP0004389/CNS1008488/CNX0953943/CNR1093075/FP200005151_L01_95_1.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data2/CNP0004389/CNS1008488/CNX0953943/CNR1093075/FP200005151_L01_95_2.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data2/CNP0004389/CNS1008489/CNX0953944/CNR1093076/DP8400026127BL_L01_70_1.fq.gz
wget -c ftp://ftp.cngb.org/pub/CNSA/data2/CNP0004389/CNS1008489/CNX0953944/CNR1093076/DP8400026127BL_L01_70_2.fq.gz
样本信息:
| sample_accession | sample_name | sample_title | dev_stage | tissue |
| CNS0809355 | RC-cDNA-0D-1 | RC-0D-c rep1 | CIM-0D | root |
| CNS0809356 | RC-cDNA-0D-2 | RC-0D-c rep2 | CIM-0D | root |
| CNS0809357 | RC-cDNA-0D-3 | RC-0D-c rep3 | CIM-0D | root |
| CNS0809358 | RC-Index-0D-1 | RC-0D-I rep1 | CIM-0D | root |
| CNS0809359 | RC-Index-0D-2 | RC-0D-I rep2 | CIM-0D | root |
| CNS0809360 | RC-Index-0D-3 | RC-0D-I rep3 | CIM-0D | root |
| CNS0809361 | RC-cDNA-1D-1 | RC-1D-c rep1 | CIM-1D | root |
| CNS0809363 | RC-Index-1D-1 | RC-1D-I rep1 | CIM-1D | root |
| CNS0809365 | RC-cDNA-4D-1 | RC-4D-c rep1 | CIM-4D | root |
| CNS0809366 | RC-cDNA-4D-2 | RC-4D-c rep2 | CIM-4D | root |
| CNS0809367 | RC-Index-4D-1 | RC-4D-I rep1 | CIM-4D | root |
| CNS0809368 | RC-Index-4D-2 | RC-4D-I rep2 | CIM-4D | root |
| CNS1008488 | RC-cDNA-1D-2 | RC-1D-c rep2 | CIM-1D | root |
| CNS1008489 | RC-Index-1D-2 | RC-1D-I rep2 | CIM-1D | root |
参考基因组建立:
# Download reference files
wget https://ftp.ensemblgenomes.ebi.ac.uk/pub/plants/release-62/fasta/arabidopsis_thaliana/dna/Arabidopsis_thaliana.TAIR10.dna.toplevel.fa.gz
wget https://ftp.ensemblgenomes.ebi.ac.uk/pub/plants/release-62/gtf/arabidopsis_thaliana/Arabidopsis_thaliana.TAIR10.62.gtf.gz
# Extract files
gzip -d Arabidopsis_thaliana.TAIR10.dna.toplevel.fa.gz
gzip -d Arabidopsis_thaliana.TAIR10.62.gtf.gz
# Build reference
dnbc4tools tools mkgtf \
--action stat \
--ingtf Arabidopsis_thaliana.TAIR10.62.gtf \
--output gtfstat.txt \
--type gene_biotype
dnbc4tools tools mkgtf --ingtf Arabidopsis_thaliana.TAIR10.62.gtf --output genes.filter.gtf --type gene_biotype \
--include protein_coding,lncRNA,lincRNA,antisense,IG_V_gene,IG_LV_gene,IG_J_gene,IG_C_gene,IG_V_pseudogene,IG_J_pseudogene,IG_C_pseudogene,TR_V_gene,TR_D_gene,TR_J_gene,TR_C_gene,rRNA
dnbc4tools rna mkref --ingtf genes.filter.gtf --fasta Arabidopsis_thaliana.TAIR10.dna.toplevel.fa --threads 10 --species Arabidopsis_thaliana
dnbc4toolsf分析:
样本有点多,单细胞转录组数据量大,只演示其中4个样本;
dnbc4tools rna run \
--name 1D1 \
--cDNAfastq1 ~/BGI-sc/data/ara/DP8400025960BR_L01_97_1.fq.gz \
--cDNAfastq2 ~/BGI-sc/data/ara/DP8400025960BR_L01_97_2.fq.gz \
--oligofastq1 ~/BGI-sc/data/ara/DP8400026128BL_L01_71_1.fq.gz \
--oligofastq2 ~/BGI-sc/data/ara/DP8400026128BL_L01_71_2.fq.gz \
--genomeDir ~/BGI-sc/ref/arabidopsis_thaliana/Arabidopsis_thaliana \
--threads 20
dnbc4tools rna run \
--name 1D2 \
--cDNAfastq1 ~/BGI-sc/data/ara/FP200005151_L01_95_1.fq.gz \
--cDNAfastq2 ~/BGI-sc/data/ara/FP200005151_L01_95_2.fq.gz \
--oligofastq1 ~/BGI-sc/data/ara/DP8400026127BL_L01_70_1.fq.gz \
--oligofastq2 ~/BGI-sc/data/ara/DP8400026127BL_L01_70_2.fq.gz \
--genomeDir ~/BGI-sc/ref/arabidopsis_thaliana/Arabidopsis_thaliana \
--threads 20
dnbc4tools rna run \
--name 4D1 \
--cDNAfastq1 ~/BGI-sc/data/ara/FP200003884_L01_27_1.fq.gz \
--cDNAfastq2 ~/BGI-sc/data/ara/FP200003884_L01_27_2.fq.gz \
--oligofastq1 ~/BGI-sc/data/ara/DP8400024433BL_L01_78_1.fq.gz \
--oligofastq2 ~/BGI-sc/data/ara/DP8400024433BL_L01_78_2.fq.gz \
--genomeDir ~/BGI-sc/ref/arabidopsis_thaliana/Arabidopsis_thaliana \
--threads 20
dnbc4tools rna run \
--name 4D2 \
--cDNAfastq1 ~/BGI-sc/data/ara/FP200003884_L01_31_1.fq.gz \
--cDNAfastq2 ~/BGI-sc/data/ara/FP200003884_L01_31_2.fq.gz \
--oligofastq1 ~/BGI-sc/data/ara/DP8400024433BL_L01_80_1.fq.gz \
--oligofastq2 ~/BGI-sc/data/ara/DP8400024433BL_L01_80_2.fq.gz \
--genomeDir ~/BGI-sc/ref/arabidopsis_thaliana/Arabidopsis_thaliana \
--threads 20
参考文献:
- Yin R, Chen R, Xia K ...A single-cell transcriptome atlas reveals the trajectory of early cell fate transition during callus induction in ArabidopsisPlant Communications, 2024; 5
- 发表于 2天前
- 阅读 ( 59 )
- 分类:转录组
你可能感兴趣的文章
- 玉米根单细胞转录组细胞类型注释代码 30 浏览
- scanpy 绘制marker基因点图 79 浏览
- 植物单细胞转录组面临的挑战 132 浏览
- 植物单细胞转录组文章 2049 浏览
- 拟南芥叶单细胞分析流程 1725 浏览
