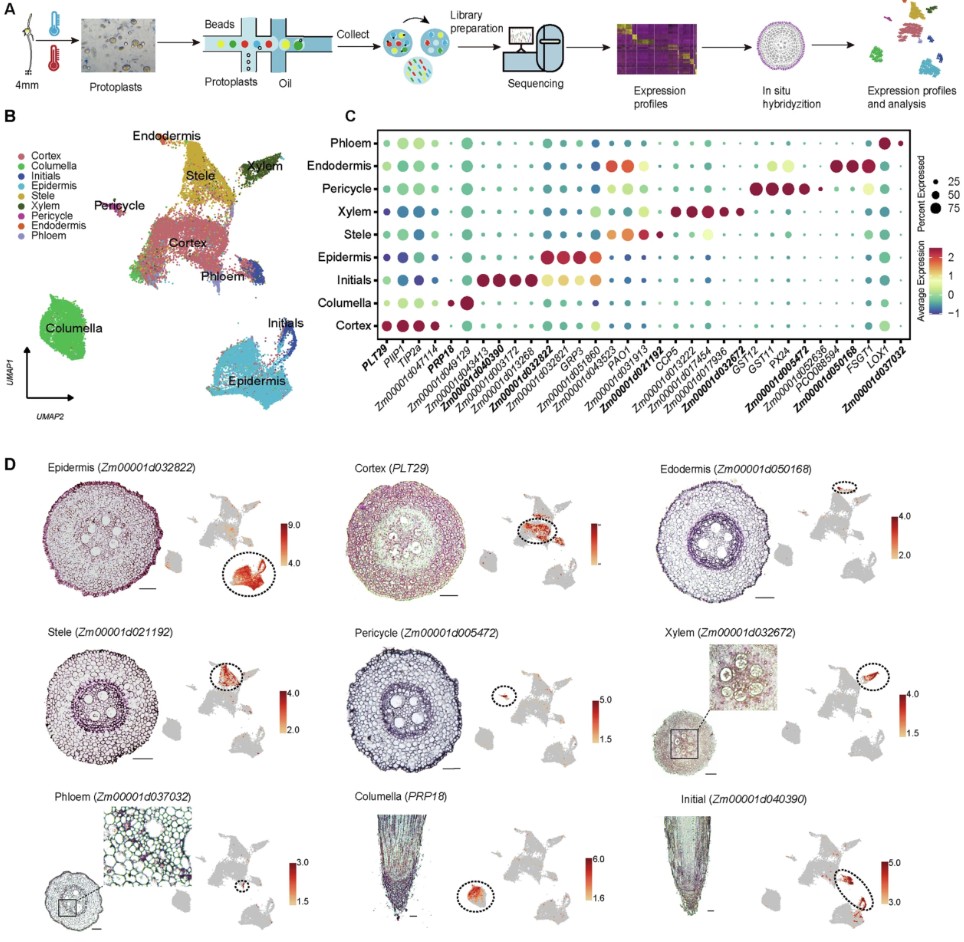
玉米根尖热处理单细胞数据分析
玉米根尖热处理单细胞数据分析
1.参考基因组下载
wget -c https://ftp.ensemblgenomes.ebi.ac.uk/pub/plants/release-62/fasta/zea_mays/dna/Zea_mays.Zm-B73-REFERENCE-NAM-5.0.dna.toplevel.fa.gz
wget -c https://ftp.ensemblgenomes.ebi.ac.uk/pub/plants/release-62/gff3/zea_mays/Zea_mays.Zm-B73-REFERENCE-NAM-5.0.62.gff3.gz
wget -c https://ftp.ensemblgenomes.ebi.ac.uk/pub/plants/release-62/gtf/zea_mays/Zea_mays.Zm-B73-REFERENCE-NAM-5.0.62.gtf.gz
2. cellranger 索引建立
#可选操作,选择蛋白编码的基因:
#cellranger mkgtf P_aphrodite_genomic_scaffold_v1.0_gene.gtf P_aphrodite_genomic_scaffold_v1.0_gene.filtered.gtf --attribute=gene_biotype:protein_coding
#10X官方参考基因组索引建立参考:https://www.10xgenomics.com/cn/support/software/space-ranger/latest/advanced/custom-references
cellranger mkref --nthreads 20 --genome=B73 --fasta=Zea_mays.Zm-B73-REFERENCE-NAM-5.0.dna.toplevel.fa --genes=Zea_mays.Zm-B73-REFERENCE-NAM-5.0.62.gtf
3.示例数据下载
wget -c ftp://download.big.ac.cn/gsa2/CRA015077/CRR1070725/CRR1070725_f1.fq.gz
wget -c ftp://download.big.ac.cn/gsa2/CRA015077/CRR1070726/CRR1070726_f1.fq.gz
wget -c ftp://download.big.ac.cn/gsa2/CRA015077/CRR1070727/CRR1070727_f1.fq.gz
wget -c ftp://download.big.ac.cn/gsa2/CRA015077/CRR1070728/CRR1070728_f1.fq.gz
wget -c ftp://download.big.ac.cn/gsa2/CRA015077/CRR1070725/CRR1070725_r2.fq.gz
wget -c ftp://download.big.ac.cn/gsa2/CRA015077/CRR1070726/CRR1070726_r2.fq.gz
wget -c ftp://download.big.ac.cn/gsa2/CRA015077/CRR1070727/CRR1070727_r2.fq.gz
wget -c ftp://download.big.ac.cn/gsa2/CRA015077/CRR1070728/CRR1070728_r2.fq.gz
4. 改名字 方便cellranger读入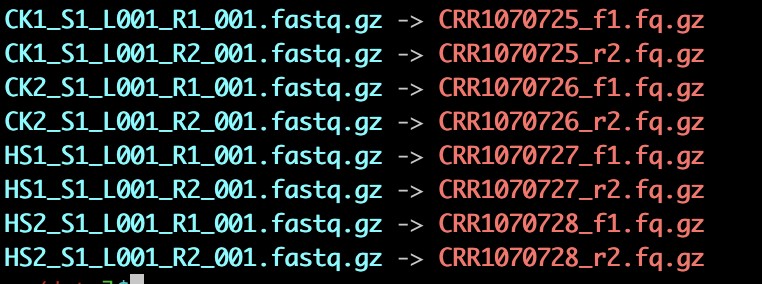

5. 准备样本信息文件 meta.tsv :
SampleID Run Accession Group CK1 PU1 CRR1070725 CK CK2 PU2 CRR1070726 CK HS1 EW01 CRR1070727 HS HS2 EW02 CRR1070728 HS
6. cellranger 分析
cat ../meta.tsv|sed '1d'|while read SampleID Run Accession Group;do
echo "Processing $SampleID"
cellranger count --id=$SampleID \
--fastqs=$datadir/ \
--sample=$SampleID \
--localcores 20 --create-bam=true \
--transcriptome=$refdir
#velocyto run10x -m $refdir/genes/mm39_rmsk.gtf ../01.cellranger/$SampleID/ $refdir/genes/genes.gtf
done
7. 质控:
cat ../meta.tsv|sed '1d'|while read SampleID Run Accession Group;do
echo "Processing $SampleID"
Rscript $scripts/seurat_sc_qc.r \
-d ../01.cellranger/$SampleID/outs/filtered_feature_bc_matrix/ \
-p $SampleID --project $SampleID \
--gene.column 1 \
--min.cells 3 \
--nGene.min 200 \
--nGene.max 9000 \
--mito.gene.pattern "^MT-" --percent_mito 10 \
--pt.gene.pattern "^PT-" --percent_pt 10 \
--ribo.gene.pattern "^RP-" --percent_ribo 10 \
--log10GenesPerUMI 0.8 \
--metadata.col.name SampleID Run Accession Group \
--metadata.value $SampleID $Run $Accession $Group
#2.3 双细胞分析
#DoubletFinder 分析需要做分群聚类分析之后才可以分析,另外作者不建议合并之后分析:
#参考:https://github.com/chris-mcginnis-ucsf/DoubletFinder?tab=readme-ov-file#input-scrna-seq-data
# 因此这里需要提前跑一下seurat的分群聚类分析
#seurat默认的标准化; 另外 SCT标准化需要加参数:--sctransform
Rscript $scripts/seurat_sc_cluster.r --rds $SampleID.afterQC.qs \
--resolution 0.5 -d 30 \
-p $SampleID -o $SampleID --cpu 10
## 如果是 SCT 标准化需要加参数:--sct
## 如果要去除双细胞增加参数:--removeDoubletCells
Rscript $scripts/DoubletFinder.r -i $SampleID/$SampleID.qs \
-p $SampleID --annotations seurat_clusters --removeDoubletCells
done
8. 聚类分群:
# #合并数据
Rscript $scripts/merge_seurat_obj.r -i ../02.data_QC/*.doubletFinder.qs -p all.sample.merged
#不整合数据直接分析:标准化,降维 , 聚类
# Rscript $scripts/seurat_sc_cluster.r --cpu 20 --rds all.sample.merged.qs \
# --umap.method "umap-learn" --n.neighbors 30 --min.dist 0.2 \
# --resolution 0.8 -d 40 --integrate.method HarmonyIntegration --batch.id SampleID \
# -p root -o root --high.variable.genes 2000 --remove_gene_regex "^RT-" \
# --skip_JackStraw_ScoreJackStraw
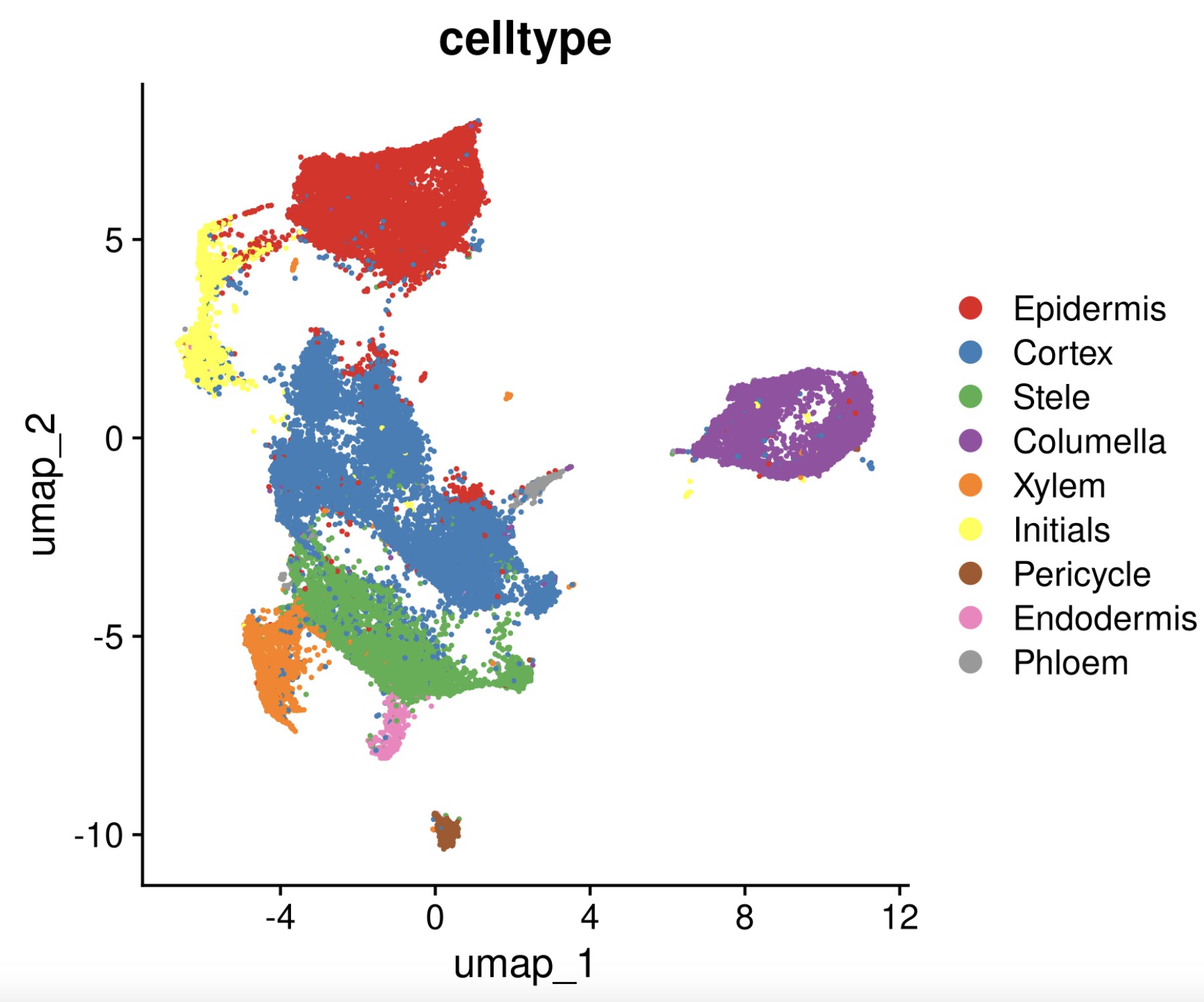
Rscript $scripts/seurat_sc_cluster.r --cpu 20 --rds all.sample.merged.qs \
--resolution 0.6 -d 30 --integrate.method HarmonyIntegration --batch.id SampleID \
-p root -o root --min.dist 0.1 --high.variable.genes 2000 --remove_gene_regex "^RT-" \
--skip_JackStraw_ScoreJackStraw
9.细胞注释:
#点图展示 marker基因 seurat_clusters
# https://www.nature.com/articles/s41467-024-55485-3
# Zm00001d012081 (PLT29)
# Zm00001d021192 (umc2686b)
# Zm00001d004089 (PRP18)
#
#
# Phloem → 韧皮部
# Endodermis → 内皮层
# Pericycle → 周鞘
# Xylem → 木质部
# Stele → 中柱(维管柱)
# Epidermis → 表皮
# Initials → 初始细胞(分生组织起始细胞)
# Columella → 柱状细胞(根冠柱状细胞)
# Cortex → 皮层
# https://www.science.org/doi/10.1126/science.abj2327#supplementary-materials
# MYB46Zm00001d038878Xylem
# LCR74Zm00001d027500Endodermis
# WOL1Zm00001d014297Stele
# TMO5Zm00001d004630Stele
# RHD6Zm00001d017804Epidermis
# ANAC074Zm00001d051140Putative pith
# CO2Zm00001d051384cortex
# CO2-likeZm00001d027291cortex
# https://onlinelibrary.wiley.com/doi/full/10.1111/pbi.14097
#Zm00001d027500flower-specific gamma-thionin1FSGT1
#Zm00001d031449lipoxygenase13ZmLOX13
#Zm00001d042541lipoxygenase1ZmLOX1
#Zm00001d035689cysteine protease5CCP5
#Zm00001d024281polyamine oxidase1PAO1
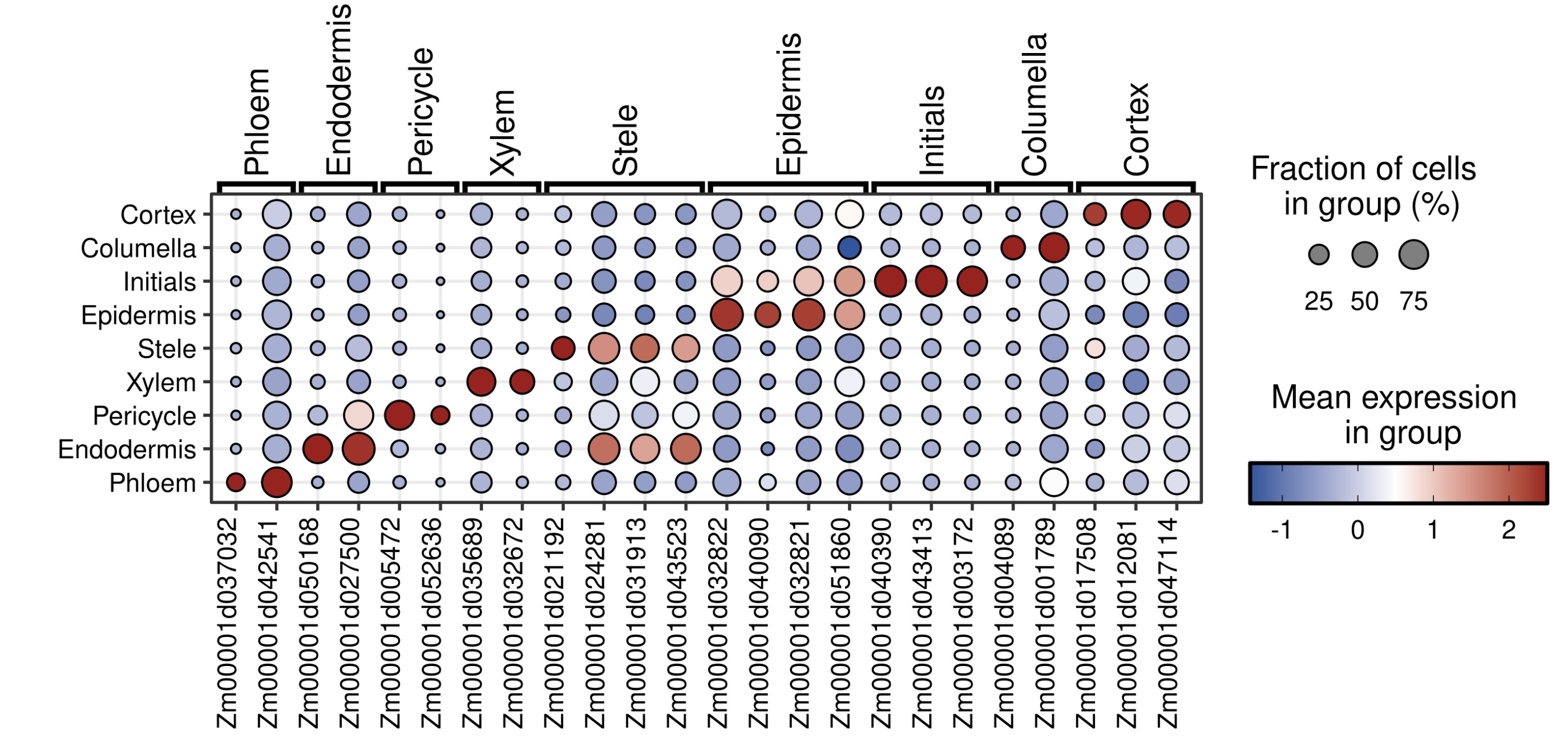
Rscript $scripts/jjDotPlot.r -i ../03.seurat_cluster/root.qs --group.by seurat_clusters \
--markerGeneCelltype.order Phloem Endodermis Pericycle Xylem Stele Epidermis Initials Columella Cortex \
--markerGene '{"Phloem": ["Zm00001d037032","Zm00001d042541"], \
"Endodermis": ["Zm00001d050168","Zm00001d027500"], \
"Pericycle": ["Zm00001d005472","Zm00001d052636"], \
"Xylem": ["Zm00001d035689", "Zm00001d032672","Zm00001d0013222","Zm00001d0017454","Zm00001d0017936"], \
"Stele": ["Zm00001d021192","Zm00001d024281","Zm00001d031913","Zm00001d043523"], \
"Epidermis": ["Zm00001d032822","Zm00001d040090","Zm00001d032821","Zm00001d051860"], \
"Initials": ["Zm00001d040390","Zm00001d043413","Zm00001d003172","Zm00001d03268"], \
"Columella": ["Zm00001d004089","Zm00001d001789"], \
"Cortex": ["Zm00001d017508", "Zm00001d012081", "Zm00001d047114"]}' \
-p Marker.seurat_clusters --plot.margin 5 1 1 1 --anno -H 8 -W 10
参考文献:
- 发表于 2026-02-06 16:30
- 阅读 ( 342 )
- 分类:转录组
你可能感兴趣的文章
相关问题
0 条评论
请先 登录 后评论